Sebastian Ahnert

Dr Sebastian Ahnert
Royal Society University Research Fellow
Fellow of King's College
Office: 518 Mott Bld
Phone: +44(0)1223 3 39852
Email: sea31 @ cam.ac.uk
Personal web site
TCM Group, Cavendish Laboratory
19 JJ Thomson Avenue,
Cambridge, CB3 0HE UK.
Research Group
Research
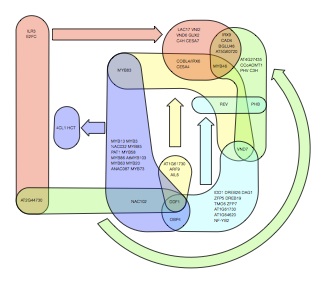
My main research interests lie on the interface of theoretical physics, biology, mathematics and computer science. I am particularly interested in using algorithmic descriptions of structures and functional systems in order to quantify and classify their complexity. Examples of the application of such approaches include pattern detection in gene expression data, the classification of protein quaternary structure and the identification of large-scale features in complex networks (see Figure). Connected to this I also have an interest in interdiscliplinary applications of network analysis, particularly in digital humanities.

In Plain English
Biology is incredibly complex, from the level of ecosystems all the way down to the level of molecules. My research tries to shed light on some of this complexity from a physicist's perspective, by applying two different tools. These are 'algorithmic information theory' and 'network analysis'. To understand the first, consider some data that a scientist has generated. Algorithmic information theory asks: "What is the shortest computer program that could output this data?" The length of such a program is one way of measuring the data's complexity. This concept can be used to compare and classify complex biological data, and to search for patterns in it. We can also use it to answer questions such as "How does complexity increase in biological evolution?". My second tool is network analysis. Interest in this area has grown rapidly over the past fifteen years because of the abundance of network data that now surrounds us (the Internet, Facebook, mobile phone networks, transport networks, power grids, neural networks, gene networks, etc.). I'm interested in identifying large-scale structures in networks (both biological and non-biological), as well as extending the reach of network analysis by applying it to history, public health, and food science.
Featured Publications
- Principles of assembly reveal a periodic table of protein complexes Science 350 6626 1331 & aaa2245 (2015) (see also University of Cambridge repository copy)
- A tractable genotype-phenotype map modelling the self-assembly of protein quaternary structure. J. Roy. Soc. Interface 11 20140249 (2014) (see also: arXiv:1311.0399v1)
- The rich club of the C. elegans neuronal connectome. J. Neurosci. 33 6380 - 6387 (2013)
- Flavor network and the principles of food pairing Scientific Reports 1 (2011) (see also: arXiv:1111.6074v1)
- Oscillating gene expression determines competence for periodic Arabidopsis root branching. Science 329 1306 - 1311 (2010)
- Self-assembly, modularity, and physical complexity. Phys. Rev. E 82 026117 (2010) (see also: arXiv:0912.3464v2)
- Unbiased pattern detection in microarray data series. Bioinformatics 22 1471 - 1476 (2006)