Research: biology
The coordination of cell proliferation and fate specification is central to specification and maintenance of tissues. In development, systems must be tightly-regulated to ensure that cells are generated in the correct number, sequence and composition whilst, in adult, a delicate balance between proliferation and differentiation is essential to ensure 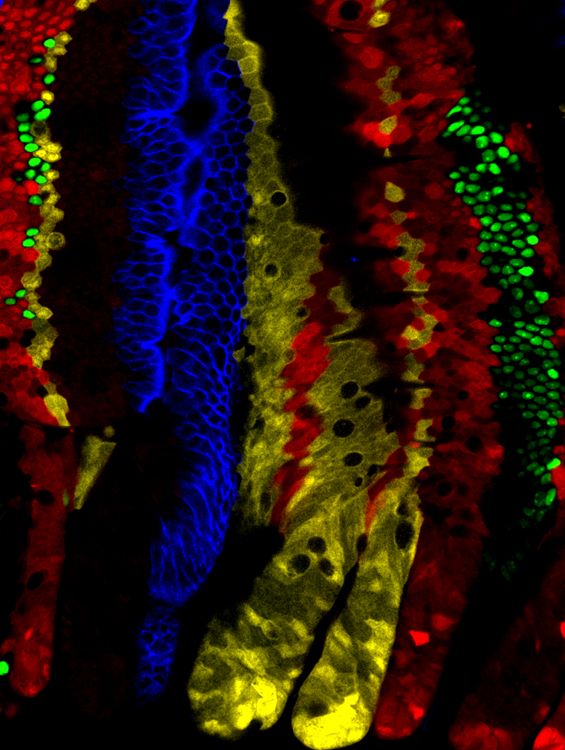 homeostasis. Through a programme of multidisciplinary and collaborative research, we target unifying principles of stem and progenitor cell regulation in the development and maintenance of tissues, and factors leading to their dysregulation in the transition to diseased states.
homeostasis. Through a programme of multidisciplinary and collaborative research, we target unifying principles of stem and progenitor cell regulation in the development and maintenance of tissues, and factors leading to their dysregulation in the transition to diseased states.
Theories of adult tissue maintenance place stem cells at the apex of proliferative hierarchies, possessing the lifetime property of self-renewal. In homeostasis the number of stem cells must remain fixed imposing an absolute requirement for fate asymmetry in the daughters of dividing cells such that only half are retained. Such fate asymmetry can be achieved either by being the invariant result of every division or by being orchestrated from the whole population, where cell fate following stem cell division is specified only up to some probability. These alternative models suggest different mechanisms of fate regulation, yet their identification in most tissues has remained elusive.
To study stem cell fate behaviour, emphasis has been placed on genetic labelling using transgenic animal models. In this approach, the activation of a fluorescent reporter gene can be used to mark a targeted cell population and their differentiating progeny. By adapting methods based on non-equilibrium statistical mechanics and population dynamics, the quantitative analysis of lineage-labelled cells -termed clones- allows the hierarchy and fate behaviour of stem and progenitor cells to be recovered. By defining the functional fate behaviour of cell populations, we are targeting the molecular mechanisms that regulate cell fate choice.
To develop this programme, we expanding our own experimental activities at the Cambridge Stem Cell Institute, while maintaining multiple collaborations with partner experimental labs, studying stem cell self-renewal in blood, brain, intestine, skin epidermis, stomach and testis. Our approach draws heavily on cell lineage tracing strategies, from in vivo live-imaging and genetic labeling of animal models to naturally occuring genomic and mitochondrial DNA mutations in human tissues. Current collaborators include Laure Bally-Cuif (zebrafish brain), Cedric Blanpain (skin development and maintenance), Hans Clevers (intestine and brain), Rakesh Heer (human prostate), Bon-Kyoung Koo (intestine and stomach), Jacco van Rheenen (intestinal maintenance and mammary gland development) and Shosei Yoshida (spermatogenesis).
Alongside research into the maintenance of adult tissues, we are interested in understanding how cells regulate the late-stage development, repair and regeneration of adult 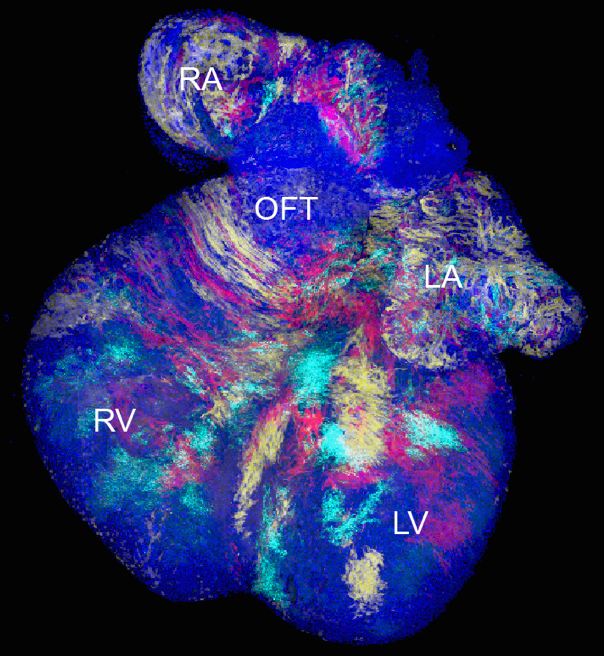 tissues. Current collaborators include Cedric Blanpain (prostate and heart), Kim Jensen (intestine and sebacious gland), Anna Philpott (pancreas) and Songhai Shi (neocortex).
tissues. Current collaborators include Cedric Blanpain (prostate and heart), Kim Jensen (intestine and sebacious gland), Anna Philpott (pancreas) and Songhai Shi (neocortex).
