

Next: Visualisation
Up: Trajectory Analysis
Previous: Trajectory Analysis
Subsections
This is available for download by clicking the following
link. mdtep-4-4.tar.gz
(releases 4.4 or later. For for compatibility with earlier releases
use mdtep-old.tar.gz).
The included makefile should be easily modified for any system.
MDTEP has been tested only on Intel Linux and Sun Solaris
platforms, but no problems on other systems are anticipated. Note
that GNUMake must be used with this makefile.
Once compiled, an MDTEP input file should be placed
in the same directory as the .md and .castep file
from which information is to be extracted. The program
should then be invoked with:
mdtep myinputfile.input
The structure of the input file uses FORTRAN name-lists. An
example is shown below.
&RunName
SeedName = 'mymdrun'
 Seedname of CASTEP calculation
Seedname of CASTEP calculation
/
&Positions
 MD step at which to begin analysis
MD step at which to begin analysis
Start = 1
 MD step at which to begin analysis
MD step at which to begin analysis
Final = 800
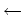 MD step at which to end analysis
MD step at which to end analysis
/
&CalcFlags
CalcRDF = .TRUE.
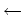 Calculate radial distribution function
Calculate radial distribution function
CalcVAC = .TRUE.
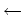 Calculate velocity autocorrelation function
Calculate velocity autocorrelation function
CalcMSD = .TRUE.
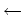 Calculate mean square displacement
Calculate mean square displacement
CalcHeatCap = .TRUE.
 Use fluctuations to calculate heat capacity
Use fluctuations to calculate heat capacity
CalcAlphaP = .TRUE.
 Use fluctuations to calculate expansion coefficient
Use fluctuations to calculate expansion coefficient
CalcBetaT = .TRUE.
 Use fluctuations to calculate bulk modulus
Use fluctuations to calculate bulk modulus
CalcTempDist = .TRUE.
 Calculate distribution of temperature samples
Calculate distribution of temperature samples
CalcVolDist = .TRUE.
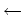 Calculate distribution of volume samples
Calculate distribution of volume samples
XmolFile = .TRUE.
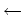 Generate <seedname>.xmol for visualisation
Generate <seedname>.xmol for visualisation
XFSFile = .TRUE.
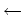 Generate <seedname>.axsf for visualisation
Generate <seedname>.axsf for visualisation
/
&VACParams
VACLength = 128
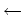 Length in MD steps over which VACF calculated
Length in MD steps over which VACF calculated
VACInterval = 1
 Steps between VACF functions to be averaged over
Steps between VACF functions to be averaged over
/
&MSDParams
MSDLength = 128
 Length in steps over which MSD is calculated
Length in steps over which MSD is calculated
MSDInterval = 1
 Steps between MSDs to be averaged over
Steps between MSDs to be averaged over
/
Notes:
- Quantities calculated from fluctuations require many thousands of MD steps to
produce accurate quantities.
- Heat capacity calculations are only valid for simulations using a thermostat.
- Expansion coefficient and bulk modulus calculations should only be performed
for variable cell MD runs.
- All calculations based on fluctuations invoke a blocking analysis to estimate
the error in the calculated quantity - see Haile [4] for details.
- The distribution of temperature samples is written to file along with the
ideal case for large N.
- The distribution of volume samples is written to file along with the ideal
case calculated assuming the estimate of
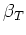 obtained is accurate.
obtained is accurate.
Each calculated quantity is written to a separate output file, prefixed
with the calculation seedname. The names of these files should be self
explanatory.
As well as computing the quantities specified in the input
file, MDTEP writes the following files, the first three
of which are suitable for direct plotting in e.g. gnu-plot
or xmgrace.
This file contains four columns of data, the first being
the elapsed simulation time in picoseconds. The subsequent
columns list the total ionic kinetic energy, the Hamiltonian
energy for the appropriate ensemble, and the value
of the Born-Oppenheimer Hamiltonian.
This file lists for each time-step (in picoseconds)
the nine components of the matrix of cell vectors in
atom units of length.
This file contains four columns, listing time in
picoseconds, temperature in Kelvin, pressure in
mega-pascals, and volume in atomic units.
This file lists the three components of the force
of each atom. The inner loop is over the number
of atoms, and the outer loop over the number
of time-steps.
This is an intermediate format used when creating
.xmol and .xsf files.


Next: Visualisation
Up: Trajectory Analysis
Previous: Trajectory Analysis
David Quigley
2005-05-10